This page is an attempt to translate into R the parts of the equivalent Stata FAQ page.
First off, let’s start with what a significant continuous by continuous interaction means. It means that the slope of one continuous variable on the response variable changes as the values on a second continuous change.
Multiple regression models often contain interaction terms. This FAQ page covers the situation in which there is a moderator variable which influences the regression of the dependent variable on an independent/predictor variable. In other words, a regression model that has a significant two-way interaction of continuous variables.
There are several approaches that one might use to explain an interaction of two continuous variables. The approach that we will demonstrate is to compute simple slopes, i.e., the slopes of the dependent variable on the independent variable when the moderator variable is held constant at different combinations of values from very low to very high.
We will consider a regression model which includes a continuous by continuous interaction of a predictor variable with a moderator variable. In the formula, Y is the response variable, X the predictor (independent) variable with Z being the moderator variable. The term XZ is the interaction of the predictor with the moderator.
Y = b0 + b1X + b2Z + b3XZ
We will illustrate the simple slopes process using the hsbdemo dataset that has a statistically significant continuous by continuous interaction when read is the response variable, math is the predictor and socst is the moderator variable. We will first look at summary statistics for all three variables.
library(foreign)
library(msm)
d <- read.dta("https://stats.idre.ucla.edu/stat/data/hsbdemo.dta")
attach(d)
summary(cbind(read, math, socst))
read math socst
Min. :28.00 Min. :33.00 Min. :26.0
1st Qu.:44.00 1st Qu.:45.00 1st Qu.:46.0
Median :50.00 Median :52.00 Median :52.0
Mean :52.23 Mean :52.65 Mean :52.4
3rd Qu.:60.00 3rd Qu.:59.00 3rd Qu.:61.0
Max. :76.00 Max. :75.00 Max. :71.0
With these value ranges in mind, we run our model using the glm command.
m1 <- glm(read ~ math*socst)
summary(m1)
Call:
glm(formula = read ~ math * socst)
Deviance Residuals:
Min 1Q Median 3Q Max
-18.6071 -4.9228 -0.7195 4.5912 21.8592
Coefficients:
Estimate Std. Error t value Pr(>|t|)
(Intercept) 37.842715 14.545210 2.602 0.00998 **
math -0.110512 0.291634 -0.379 0.70514
socst -0.220044 0.271754 -0.810 0.41908
math:socst 0.011281 0.005229 2.157 0.03221 *
---
Signif. codes: 0 "***" 0.001 "**" 0.01 "*" 0.05 "." 0.1 " " 1
(Dispersion parameter for gaussian family taken to be 48.44213)
Null deviance: 20919.4 on 199 degrees of freedom
Residual deviance: 9494.7 on 196 degrees of freedom
AIC: 1349.6
Number of Fisher Scoring iterations: 2
Please note that the interaction, math:socst, is statistically significant with a p-value of 0.03221.
Next, we compute the slope for read on math while holding the value of the moderator variable, socst, constant at values running from 30 to 75. To do this, we will find the total coefficient for math in the model equation for each value of socst. Using the equation presented in the introduction and allowing math to be X and socst to be Z, we can see that the total coefficient for math is b1 + b3*socst. Below, we go through this logic in R.
m1$coef (Intercept) math socst math:socst 37.84271468 -0.11051227 -0.22004419 0.01128072 at.socst <- seq(30, 75, 5) slopes <- m1$coef[2] + m1$coef[4]*at.socst slopes [1] 0.2279094 0.2843130 0.3407166 0.3971202 0.4535238 0.5099274 0.5663311 [8] 0.6227347 0.6791383 0.7355419
Next, we will use the delta method to estimate the standard errors of these slopes. The deltamethod command appears in the msm package. After calculating the standard errors, we find 95% confidence intervals.
estmean<-coef(m1)
var<-vcov(m1)
SEs <- rep(NA, length(at.socst))
for (i in 1:length(at.socst)){
j <- at.socst[i]
SEs[i] <- deltamethod (~ (x2) + (x4)*j, estmean, var)
}
upper <- slopes + 1.96*SEs
lower <- slopes - 1.96*SEs
cbind(at.socst, slopes, upper, lower)
at.socst slopes upper lower
[1,] 30 0.2279094 0.5071945 -0.05137570
[2,] 35 0.2843130 0.5186840 0.04994197
[3,] 40 0.3407166 0.5333616 0.14807164
[4,] 45 0.3971202 0.5537953 0.24044513
[5,] 50 0.4535238 0.5848051 0.32224254
[6,] 55 0.5099274 0.6331150 0.38673993
[7,] 60 0.5663311 0.7018605 0.43080157
[8,] 65 0.6227347 0.7864845 0.45898483
[9,] 70 0.6791383 0.8804154 0.47786117
[10,] 75 0.7355419 0.9793935 0.49169023
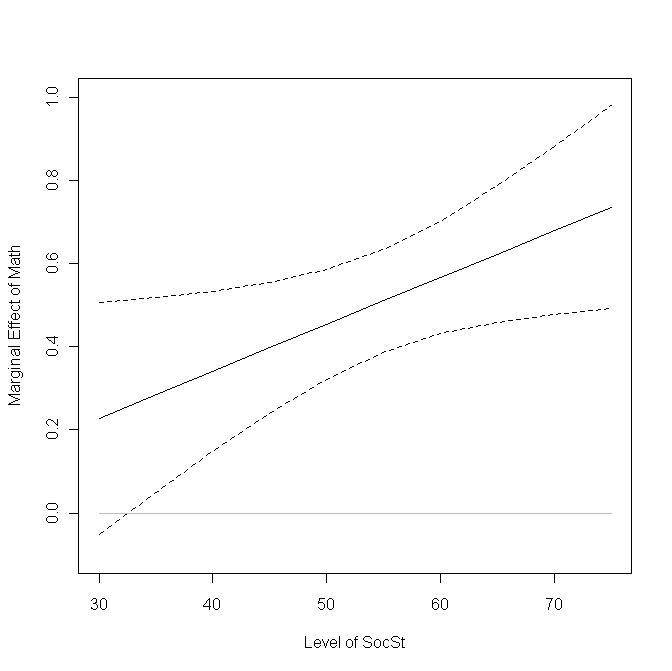
We can plot this information to show how the slope of math changes with the level of socst and where the slope is significantly different from 0.
plot(at.socst, slopes, type = "l", lty = 1, ylim = c(-.1, 1), xlab = "Level of SocSt", ylab = "Marginal Effect of Math") points(at.socst, upper, type = "l", lty = 2) points(at.socst, lower, type = "l", lty = 2) points(at.socst, rep(0, length(at.socst)), type = "l", col = "gray")