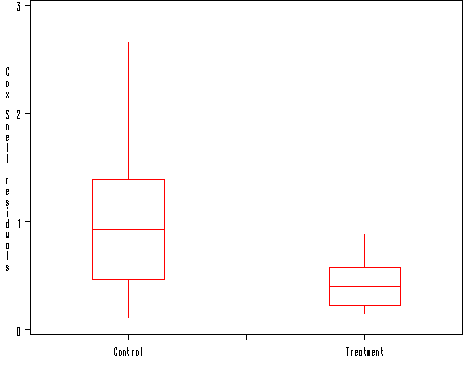Table 10.2 on page 178.
data table10_1; length group $9; input group $ censor time; datalines; control 1 1 control 1 1 control 1 2 control 1 2 control 1 3 control 1 4 control 1 4 control 1 5 control 1 5 control 1 8 control 1 8 control 1 8 control 1 8 control 1 11 control 1 11 control 1 12 control 1 12 control 1 15 control 1 17 control 1 22 control 1 23 treatment 1 6 treatment 1 6 treatment 1 6 treatment 0 6 treatment 1 7 treatment 0 9 treatment 1 10 treatment 0 10 treatment 0 11 treatment 1 13 treatment 1 16 treatment 0 17 treatment 0 19 treatment 0 20 treatment 1 22 treatment 1 23 treatment 0 25 treatment 0 32 treatment 0 32 treatment 0 34 treatment 0 35 ; run; proc lifetest data = table10_1; time time*censor(0); strata group; ods output ProductLimitEstimates = table10_2; run; proc print data = table10_2; where group = 'treatment'; run;
Obs STRATUM group time Censor Survival Failure StdErr Failed Left 23 2 treatment 0.0000 0 1.0000 0 0 0 21 24 2 treatment 6.0000 0 . . . 1 20 25 2 treatment 6.0000 0 . . . 2 19 26 2 treatment 6.0000 0 0.8571 0.1429 0.0764 3 18 27 2 treatment 6.0000 1 . . . 3 17 28 2 treatment 7.0000 0 0.8067 0.1933 0.0869 4 16 29 2 treatment 9.0000 1 . . . 4 15 30 2 treatment 10.0000 0 0.7529 0.2471 0.0963 5 14 31 2 treatment 10.0000 1 . . . 5 13 32 2 treatment 11.0000 1 . . . 5 12 33 2 treatment 13.0000 0 0.6902 0.3098 0.1068 6 11 34 2 treatment 16.0000 0 0.6275 0.3725 0.1141 7 10 35 2 treatment 17.0000 1 . . . 7 9 36 2 treatment 19.0000 1 . . . 7 8 37 2 treatment 20.0000 1 . . . 7 7 38 2 treatment 22.0000 0 0.5378 0.4622 0.1282 8 6 39 2 treatment 23.0000 0 0.4482 0.5518 0.1346 9 5 40 2 treatment 25.0000 1 . . . 9 4 41 2 treatment 32.0000 1 . . . 9 3 42 2 treatment 32.0000 1 . . . 9 2 43 2 treatment 34.0000 1 . . . 9 1 44 2 treatment 35.0000 1 . . . 9 0
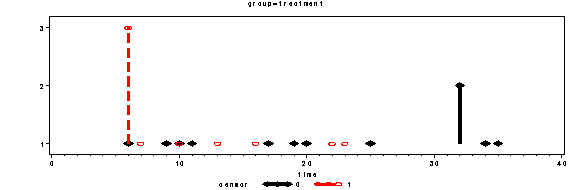
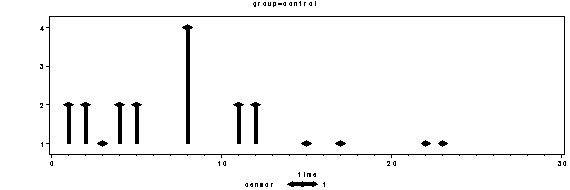
Figure 10.2 on page 179. Up to version 8.2 SAS does not have built-in dot-plots. We can get a similar plot that conveys the same type of information by using proc gplot.
proc freq data = table10_1;
tables group*censor*time / list nocum nopercent;
ods output list = mylist (keep= group censor time frequency);
run;
goptions reset = all; hsize = 6 vsize = 2;
axis1 offset = (1,1) minor = none label=(' ');
symbol1 i = needle value = P font = marker h = .5 c=black w=3;
symbol2 i = needle value = circle c = red w = 2 l=4;
proc gplot data = mylist;
by group;
plot frequency*time = censor /vaxis= axis1;
run;
quit;
Figure 10.2 on page 179.
proc lifetest data = table10_1 plot=(s); time time*censor(0); strata group; run;
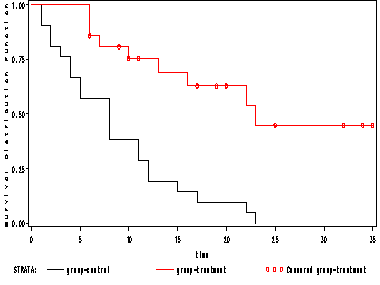
Figure 10. 4 on page 180.
goptions reset = all; proc lifetest data = table10_1 plot=(loglogs); time time*censor(0); strata group; run;
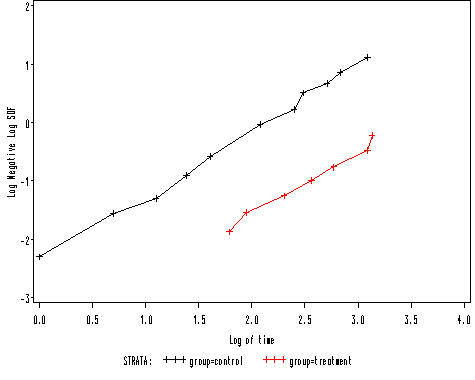
Table 10.3 on page 185.
data table10_1b;
set table10_1;
if group = 'treatment' then gp = 1;
else gp = 0;
run;
proc reliability data = table10_1b;
distribution exponential;
model time*censor(0) = gp ;
run;
Exponential Parameter Estimates
Asymptotic Normal
Standard 95% Confidence Limits
Parameter Estimate Error Lower Upper
Intercept 2.1595 0.2182 1.7318 2.5872
gp 1.5266 0.3984 0.7457 2.3075
Shape 1.0000 0.0000 1.0000 1.0000
proc reliability data = table10_1b;
distribution weibull;
model time*censor(0) = gp ;
run;
Weibull Parameter Estimates
Asymptotic Normal
Standard 95% Confidence Limits
Parameter Estimate Error Lower Upper
Intercept 2.2484 0.1660 1.9231 2.5737
gp 1.2673 0.3106 0.6585 1.8762
EV Scale 0.7322 0.1078 0.5486 0.9772
Weibull Shape 1.3658 0.2012 1.0233 1.8228
Figure 10.5 on page 186. In SAS, we can either use proc lifereg or proc reliability to perform parametric survival regression analyses. We used proc reliability here to generate different types of residuals. The Cox-Snell residual produced by proc reliability is the modified version adjusting for the right-censored observations. In the book, the unadjusted Cox-Snell residuals are plotted. So we manually generate the unadjusted Cox-Snell residuals in a data step.
ods output modobstats = residual;
proc reliability data = table10_1b;
distribution exponential;
model time*censor(0) = gp /obstats(surv dresid );
run;
data residual;
set residual;
gresid = -log(surv);
run;
axis1 order = (0 to 3 by 1) minor = none label=('Cox Snell residuals');
axis2 value=(tick=1 'Control' tick = 2 ' ' tick=3 'Treatment') label = (' ') minor = none;
proc boxplot data = residual;
plot gresid*gp =' '/vaxis=axis1 haxis=axis2 boxwidth=15 noserifs;
run;
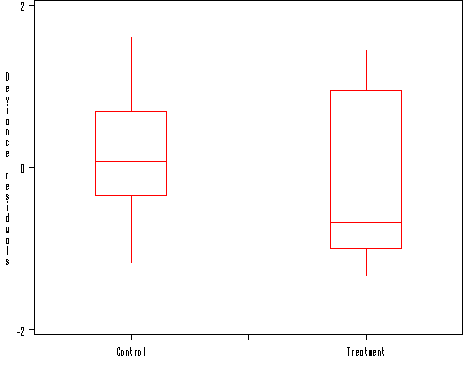
axis1 order = (-2 to 2 by 2) minor = none label=('Deviance residuals');
axis2 value=(tick=1 'Control' tick = 2 ' ' tick=3 'Treatment') label = (' ') minor = none;
proc boxplot data = residual;
plot dresid*gp =' '/vaxis=axis1 haxis=axis2 boxwidth=15 noserifs;
run;