The material in this chapter is based on the M215 (Fall 2001) Final Project Report by Dong Ding, Weihua Huang, Luohua Jiang, Liyuan Xiong and Kefei Zhou. We thank them for permission to adapt and distribute this page via our web site. This chapter is all done using the SAS macro program for the additive hazard model provided from the author’s website. This page was revised on May 27, 2010.
Section 12.2: Least-Squares Estimation for the Nonparametric, Additive Hazard Model
Example 10.1 on page 309 using the breast cancer data set described in Section 1.5. We have created a data step for it which can be downloaded here.
%include 'c:sasmacrosadditive.sas';
proc sort data = sec1_5;
by time;
run;
data bcnew;
set sec1_5;
positive = (group=2);
drop group;
run;
proc iml;
option={y, n, y, n, n};
contr={1 -1};
effects={'positive'};
timeunit={'month'};
%additive (bcnew,0.05, timeunit, effects, option, contr, betab, testb);
quit;
proc print data = betab (obs=10) noobs; run;
COL1 COL2 COL3 COL4 COL5 COL6 COL7 COL8 COL9
19 0.02778 -0.02778 0.027778 0.02778 -0.026666 -0.08222 0.08222 0.02667
22 0.02778 0.08333 0.027778 0.11453 -0.026666 -0.14114 0.08222 0.30781
23 0.02778 0.20833 0.027778 0.16954 -0.026666 -0.12395 0.08222 0.54062
25 0.05635 0.17976 0.039849 0.17193 -0.021753 -0.15721 0.13445 0.51673
30 0.08576 0.15035 0.049528 0.17442 -0.011311 -0.19151 0.18283 0.49221
34 0.11606 0.12005 0.058063 0.17704 0.002264 -0.22694 0.22986 0.46703
37 0.14731 0.08880 0.065938 0.17977 0.018078 -0.26355 0.27655 0.44115
38 0.14731 0.23165 0.065938 0.22962 0.018078 -0.21840 0.27655 0.68171
42 0.14731 0.39832 0.065938 0.28373 0.018078 -0.15779 0.27655 0.95443
46 0.17957 0.36606 0.073406 0.28556 0.035699 -0.19363 0.32344 0.92575
Notice:
Col1: time
Col2: beta_0
col3: beta_1
col4: standard error for beta_0
col5: standard error for beta_1
col6: lower confidence limit for beta_0
col7: lower confidence limit for beta_1
col8: upper confidence limit for beta_0
col9: upper confidence limit for beta_1
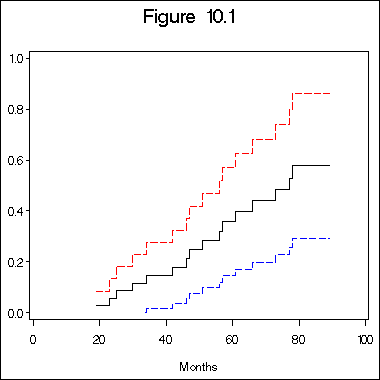
title "Figure 10.1";
axis1 label=(j=c 'Months') order=(0 to 100 by 20) minor = none;
axis2 label=(a=90 j=c 'Estimated Baseline Cumulative Hazard Rate for Immunoperoxidase Negative Patients')
order=(0.0 to 1.0 by 0.2) minor = none;
symbol1 interpol=stepjr c=black l=1 value=none;
symbol2 interpol=stepjr c=blue l=3 value=none;
symbol3 interpol=stepjr c=red l=3 value=none;
proc gplot data=betab;
plot col2*col1 col6*col1 col8*col1/overlay haxis=axis1 vaxis=axis2;
run;
quit;
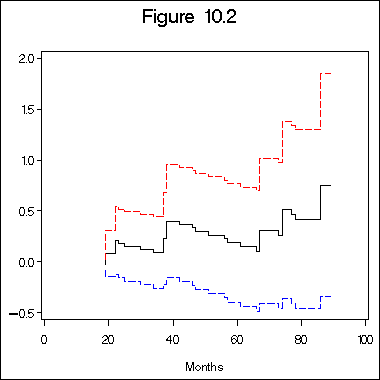
title "Figure 10.2";
axis1 label=(j=c 'Months') order=(0 to 100 by 20) minor=none;
axis2 label=(a=90 j=c 'Estimated Cumulative Excess Risk for Immunoperosidase Positive Patients')
order=(-0.5 to 2.0 by 0.5) minor=none;
proc gplot data=betab;
plot col3*col1 col7*col1 col9*col1/overlay haxis=axis1 vaxis=axis2;
run;
quit;
title;
Example 10.2 on page 311 using data set larynx described in section 1.8.
data larynx1;
length time death z1-z3 age 8.;
set larynx;
z1=(stage=2);
z2=(stage=3);
z3=(stage=4);
age=age-64.11;
keep time death z1 z2 z3 age;
run;
proc sort data = larynx1;
by time;
run;
proc iml;
option={y, n, y, n, n};
contrast={0 1 -1 0 0, 0 0 1 -1 0};
effects={'z1', 'z2' ,'z3', 'age'};
timeunit={'Years'};
%additive (larynx1, 0.05, timeunit, effects, option, contrast, beta, test);
quit;
The data set beta created above has many columns. It has the following pattern.
Col1: time Col2-6: beta_0-beta_4 col7-11: standard errors for beta_0-beta_4 col2-16: lower confidence limit for beta_0-beta_4 col7-21: upper confidence limit for beta_0-beta_4
proc contents data = beta; run;
Alphabetic List of Variables and Attributes # Variable Type Len 1 COL1 Num 8 2 COL2 Num 8 3 COL3 Num 8 4 COL4 Num 8 5 COL5 Num 8 6 COL6 Num 8 7 COL7 Num 8 8 COL8 Num 8 9 COL9 Num 8 10 COL10 Num 8 11 COL11 Num 8 12 COL12 Num 8 13 COL13 Num 8 14 COL14 Num 8 15 COL15 Num 8 16 COL16 Num 8 17 COL17 Num 8 18 COL18 Num 8 19 COL19 Num 8 20 COL20 Num 8 21 COL21 Num 8
proc print data = beta (obs = 15); run;
Obs COL1 COL2 COL3 COL4 COL5 COL6 COL7 1 0.1 0.000015 0.000130 -0.00007 0.07751 -.000202225 0.000015 2 0.2 -0.000134 0.057630 0.00068 0.07118 0.001860612 0.000149 3 0.3 -0.000105 0.057359 0.07461 0.15573 0.001464296 0.000194 4 0.3 -0.000105 0.057359 0.07461 0.15573 0.001464296 0.000194 5 0.3 -0.000105 0.057359 0.07461 0.15573 0.001464296 0.000194 6 0.4 -0.000174 0.058012 0.07467 0.24403 0.002421188 0.000206 7 0.5 -0.000120 0.057496 0.11462 0.24540 0.001664510 0.000213 8 0.6 0.030085 0.028127 0.08400 0.21261 0.003034450 0.030205 9 0.7 0.030607 0.028573 0.12466 0.20909 0.004621786 0.030210 10 0.8 0.031678 0.029490 0.16813 0.40186 0.007880698 0.030221 11 0.8 0.031678 0.029490 0.16813 0.40186 0.007880698 0.030221 12 0.8 0.031678 0.029490 0.16813 0.40186 0.007880698 0.030221 13 1.0 0.030218 0.028240 0.20991 0.53061 0.003437778 0.030239 14 1.0 0.030218 0.028240 0.20991 0.53061 0.003437778 0.030239 15 1.3 0.060846 -0.003541 0.22597 0.50733 0.001547087 0.043152 Obs COL8 COL9 COL10 COL11 COL12 COL13 COL14 1 0.000130 0.00007 0.07751 .000202225 -0.000014 -0.00012 -0.000220 2 0.057500 0.00076 0.07777 .002072725 -0.000425 -0.05507 -0.000808 3 0.057512 0.05219 0.11324 .002696663 -0.000485 -0.05536 -0.027669 4 0.057512 0.05219 0.11324 .002696663 -0.000485 -0.05536 -0.027669 5 0.057512 0.05219 0.11324 .002696663 -0.000485 -0.05536 -0.027669 6 0.057516 0.05219 0.14360 .002861404 -0.000577 -0.05472 -0.027610 7 0.057518 0.06572 0.14361 .002959762 -0.000536 -0.05524 -0.014191 8 0.064582 0.07251 0.14730 .003261431 -0.029116 -0.09845 -0.058114 9 0.064584 0.08313 0.14735 .003627198 -0.028603 -0.09801 -0.038273 10 0.064587 0.09381 0.20297 .004369147 -0.027553 -0.09710 -0.015736 11 0.064587 0.09381 0.20297 .004369147 -0.027553 -0.09710 -0.015736 12 0.064587 0.09381 0.20297 .004369147 -0.027553 -0.09710 -0.015736 13 0.064594 0.10368 0.23962 .005441764 -0.029050 -0.09836 0.006700 14 0.064594 0.10368 0.23962 .005441764 -0.029050 -0.09836 0.006700 15 0.071930 0.11833 0.24096 .005642897 -0.023731 -0.14452 -0.005941 Obs COL15 COL16 COL17 COL18 COL19 COL20 COL21 1 -0.074405 -.000598578 0.00004 0.00038 0.00007 0.22942 0.000194 2 -0.081240 -.002201855 0.00016 0.17033 0.00217 0.22360 0.005923 3 -0.066225 -.003821066 0.00027 0.17008 0.17689 0.37768 0.006750 4 -0.066225 -.003821066 0.00027 0.17008 0.17689 0.37768 0.006750 5 -0.066225 -.003821066 0.00027 0.17008 0.17689 0.37768 0.006750 6 -0.037423 -.003187060 0.00023 0.17074 0.17695 0.52548 0.008029 7 -0.036060 -.004136518 0.00030 0.17023 0.24344 0.52687 0.007466 8 -0.076099 -.003357836 0.08929 0.15471 0.22611 0.50132 0.009427 9 -0.079704 -.002487391 0.08982 0.15516 0.28759 0.49788 0.011731 10 0.004050 -.000682672 0.09091 0.15608 0.35200 0.79966 0.016444 11 0.004050 -.000682672 0.09091 0.15608 0.35200 0.79966 0.016444 12 0.004050 -.000682672 0.09091 0.15608 0.35200 0.79966 0.016444 13 0.060962 -.007227883 0.08949 0.15484 0.41313 1.00025 0.014103 14 0.060962 -.007227883 0.08949 0.15484 0.41313 1.00025 0.014103 15 0.035066 -.009512789 0.14542 0.13744 0.45789 0.97960 0.012607
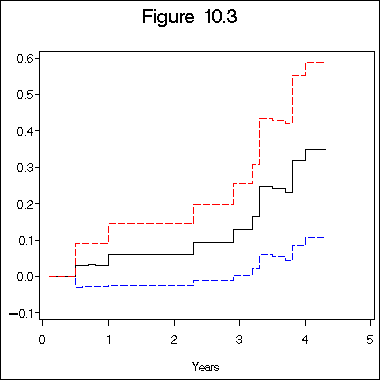
title "Figure 10.3";
axis1 label=(j=c 'Years') minor=none;
axis2 label=(a=90 j=c 'Estimated Baseline Cumulative Hazard Rate for Stage I Patients')
minor = none;
proc gplot data=beta;
plot col2*col1 col12*col1 col17*col1/overlay haxis=axis1 vaxis=axis2;
run;
quit;
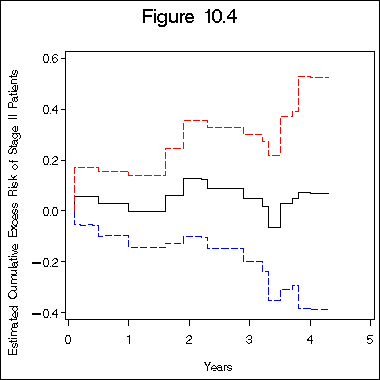
title "Figure 10.4";
axis1 label=(j=c 'Years') minor=none;
axis2 label=(a=90 j=c 'Estimated Cumulative Excess Risk of Stage II Patients')
order=(-0.4 to 0.6 by 0.2) minor = none;
proc gplot data=beta;
plot col3*col1 col13*col1 col18*col1/overlay haxis=axis1 vaxis=axis2;
run;
quit;
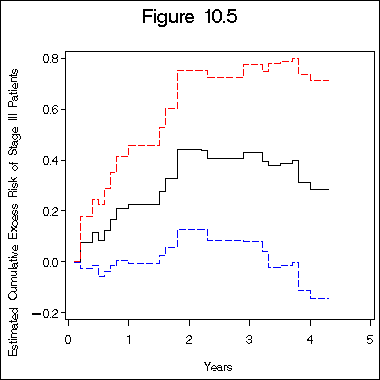
title "Figure 10.5";
axis1 label=(j=c 'Years') minor=none;
axis2 label=(a=90 j=c 'Estimated Cumulative Excess Risk of Stage III Patients')
order=(-0.2 to 0.8 by 0.2) minor=none;
proc gplot data=beta;
plot col4*col1 col14*col1 col19*col1/overlay haxis=axis1 vaxis=axis2;
run;
quit;
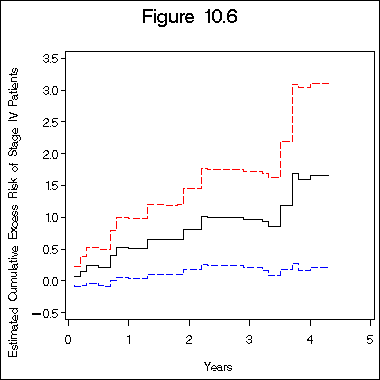
title "Figure 10.6";
axis1 label=(j=c 'Years') minor = none;
axis2 label=(a=90 j=c 'Estimated Cumulative Excess Risk of Stage IV Patients')
order=(-0.5 to 3.5 by 0.5) minor = none;
proc gplot data=beta;
plot col5*col1 col15*col1 col20*col1/overlay haxis=axis1 vaxis=axis2;
run;
quit;
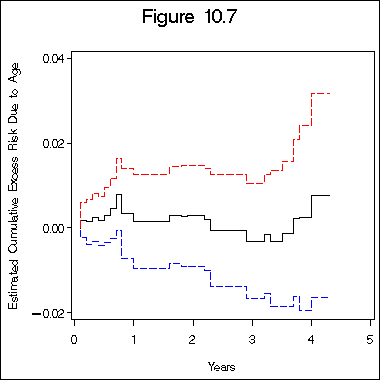
title "Figure 10.7";
axis1 label=(j=c 'Years') minor = none;
axis2 label=(a=90 j=c 'Estimated Cumulative Excess Risk Due to Age')
order=(-0.02 to 0.04 by 0.02) minor = none;
proc gplot data=beta;
plot col6*col1 col16*col1 col21*col1/overlay haxis=axis1 vaxis=axis2;
run;
quit;
title;
Section 10.3: Testing in the Nonparametric Additive Hazard Model
Table in the middle of page 323 of Example 10.2 (continued).
options nocenter;
proc iml;
option={ n, y, n, n, y};
contrast={0 1 -1 0 0, 0 0 1 -1 0};
effects={'z1', 'z2' ,'z3', 'age'};
timeunit={'Years'};
%additive (larynx1, 0.05, timeunit, effects, option, contrast, beta, test);
quit;
Additive Hazards Model
No missing data: all observations were used in analysis.
90 observations used.
Estimates are restricted to the time interval 0 to 4.30
Global Test
Chi-Square d.f p-value
10.9613 4 0.0270
Analysis of Variance
Effect Chi-Square d.f p-value
z1 0.1456 1 0.7027
z2 3.0062 1 0.0829
z3 8.4655 1 0.0036
age 0.2333 1 0.6291
Test of Linear Combinations
Contrast Matrix 0 1 -1 0 0
0 0 1 -1 0
Chi-Square d.f p-value
6.8131 2 0.0332
title "beta_1=beta_2";
proc iml;
option={ n, y, n, n, y};
contrast={0 1 -1 0 0};
effects={'z1', 'z2' ,'z3', 'age'};
timeunit={'Years'};
%additive (larynx1, 0.05, timeunit, effects, option, contrast, beta, test);
quit;
Contrast Matrix 0 1 -1 0 0
Chi-Square d.f p-value
1.5884 1 0.2076
title "beta_1=beta_3";
proc iml;
option={ n, y, n, n, y};
contrast={0 1 0 -1 0};
effects={'z1', 'z2' ,'z3', 'age'};
timeunit={'Years'};
%additive (larynx1, 0.05, timeunit, effects, option, contrast, beta, test);
quit;
Contrast Matrix 0 1 0 -1 0
Chi-Square d.f p-value
6.9390 1 0.0084
title "beta_2=beta_3";
proc iml;
option={ n, y, n, n, y};
contrast={0 0 1 -1 0};
effects={'z1', 'z2' ,'z3', 'age'};
timeunit={'Years'};
%additive (larynx1, 0.05, timeunit, effects, option, contrast, beta, test);
quit;
Contrast Matrix 0 0 1 -1 0
Chi-Square d.f p-value
3.4238 1 0.0643
